feature request #8357
closed
implement client adapter for diatom base
Added by Andreas Kohlbecker almost 5 years ago.
Updated about 2 years ago.
Description
example requests:
http --timeout 600 :8081/search 'providers==worms[diatombase]' 'query==Eunotia can' searchMode==scientificNameLike | jq '.query[] | .response[] | .taxon.taxonName.scientificName'
Files
Diatom base is fully integrated into WoRMS served throgh the same system.
The search form for DiatomBase uses in the background the same API as WoRMS with a specific preset. The following search settings in WoRMS provide the exact same reults as a search in DiatomBase (tested with the name "Navicula"):
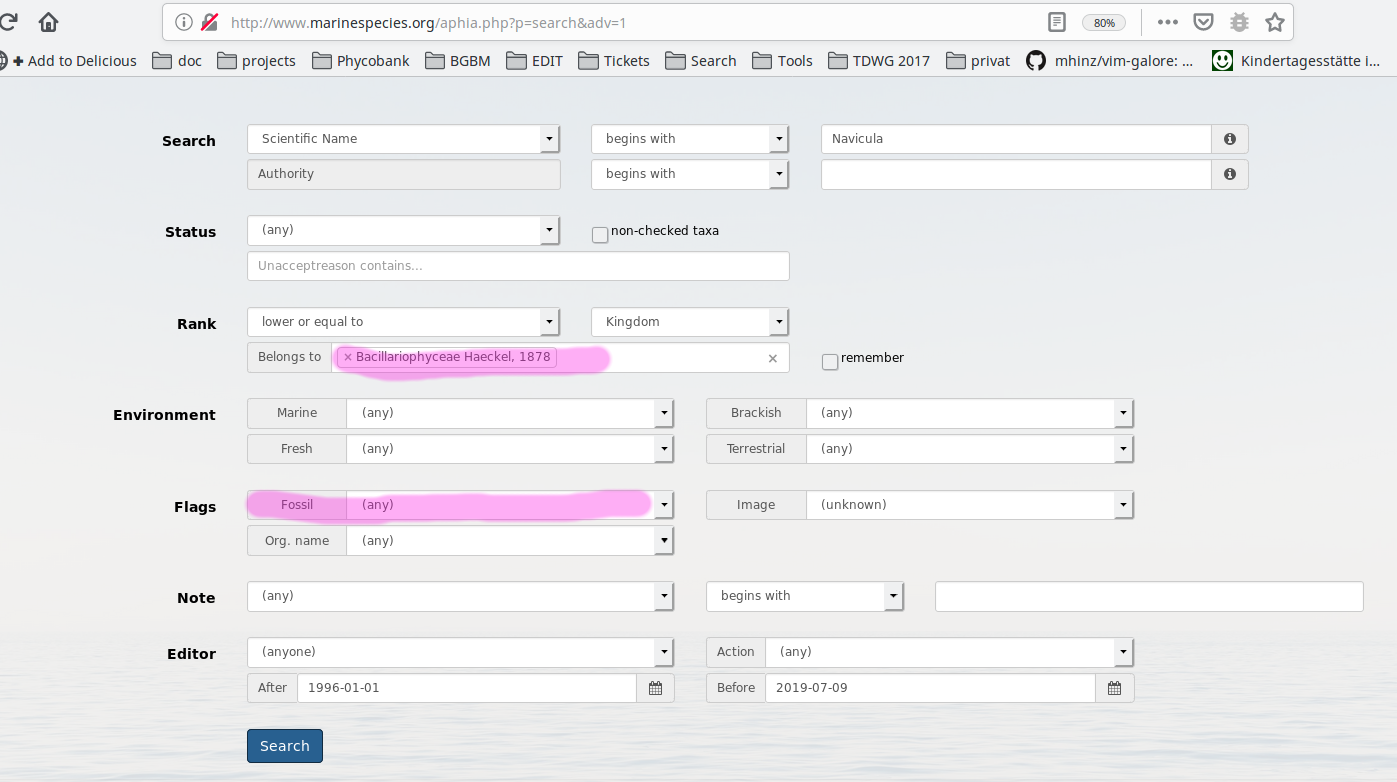
- Description updated (diff)
Bart Vanhoorne wote:
diatombase.org (and all other subregisters in WoRMS/Aphia http://www.marinespecies.org/subregisters.php) use a system for tagging taxa/species.
For Diatombase, this of course boils down to "limit to Bacillariophyceae, but there might be exceptions to this.
Conclusion, if you use the access point on diatombase.org (http://www.diatombase.org/aphia.php?p=webservice) you will always have the correctly tagged taxa, and don't need to apply a higher taxon as filter.
- Status changed from New to In Progress
- Priority changed from New to Highest
- Status changed from In Progress to Closed
- % Done changed from 0 to 100
- Target version set to 280
- Target version changed from 280 to UTIS 1.3
- Project changed from UTIS to EDIT
- Category changed from utis to utis
- Severity set to normal
Also available in: Atom
PDF
